A DIVERTICULAR DISEASE PREDICTION MODEL USING A POLYGENIC RISK SCORE
Henry D. Schaeffer*, Diane T. Smelser, H. Shanker Rao, Jeremy S. Haley, Kevin Long, Sasha Slipak, David Carey, Rebecca Hoffman
General Surgery, Geisinger Medical Center, Danville, PA
Background
Despite its prevalence and associated morbidity, we remain largely unable to predict which patients are at risk of diverticular disease (DD). To date, several clinical and genetic factors have been proposed, but no prediction model exists. Our aim was to create a prediction model for DD using a novel polygenic risk score.
Methods
A case-control cohort of patients of European ancestry who were enrolled in the MyCode Community Health Initiative biobanking program were used to evaluate the predictive ability of a PRS for DD. Using the associated electronic health record data, ICD codes for both diverticulosis and diverticulitis were used to identify patients with diverticular disease, while the remaining patients were included as controls. This patient cohort was used to evaluate the PRS developed based off the summary statistics from previously published genome-wide association studies.
Results
A total of 131,943 patients were included, of which 16,394 (12%) had DD (8,105 with diverticulitis and 8,289 with diverticulosis). The polygenic risk score for DD included 916 single nucleotide variants and had an R2 of 0.0089. There was an approximately 2.5-fold difference in disease risk between lowest and highest percentiles. Risk scores that were broken down by disease severity had R2 values of 0.0069 (diverticulosis) and 0.0093 (diverticulitis).
Conclusion
This novel predictive model provides a mechanism to stratify patients into high and low risk groups based on genetic factors and introduces the possibility of using disease risk to guide clinical decision making.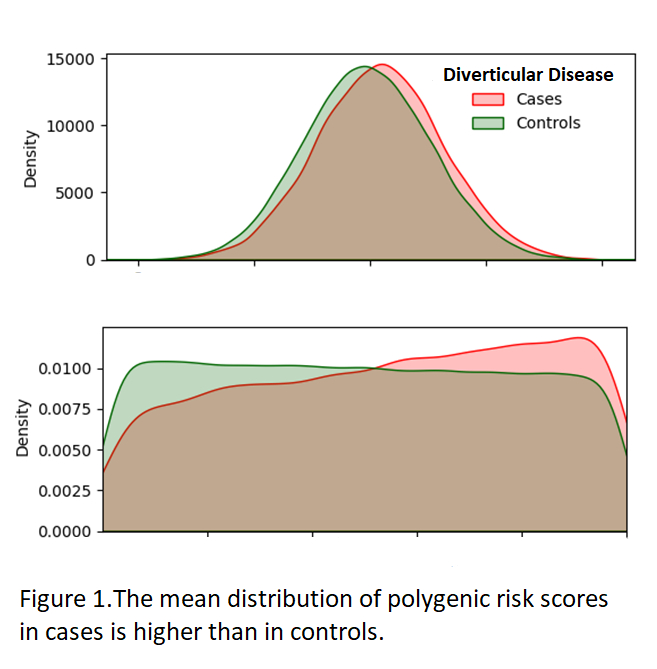

Back to 2022 Abstracts
